Note
Go to the end to download the full example code.
MixedNB Equivalence with BernoulliNB#
This example demonstrates that skbn.mixed_nb.MixedNB
produces identical results to sklearn.naive_bayes.BernoulliNB
when all features are binary (Bernoulli).
The plot shows the decision boundaries for both classifiers. As expected, the boundaries are identical, and the predicted probabilities for the dataset are all-close.
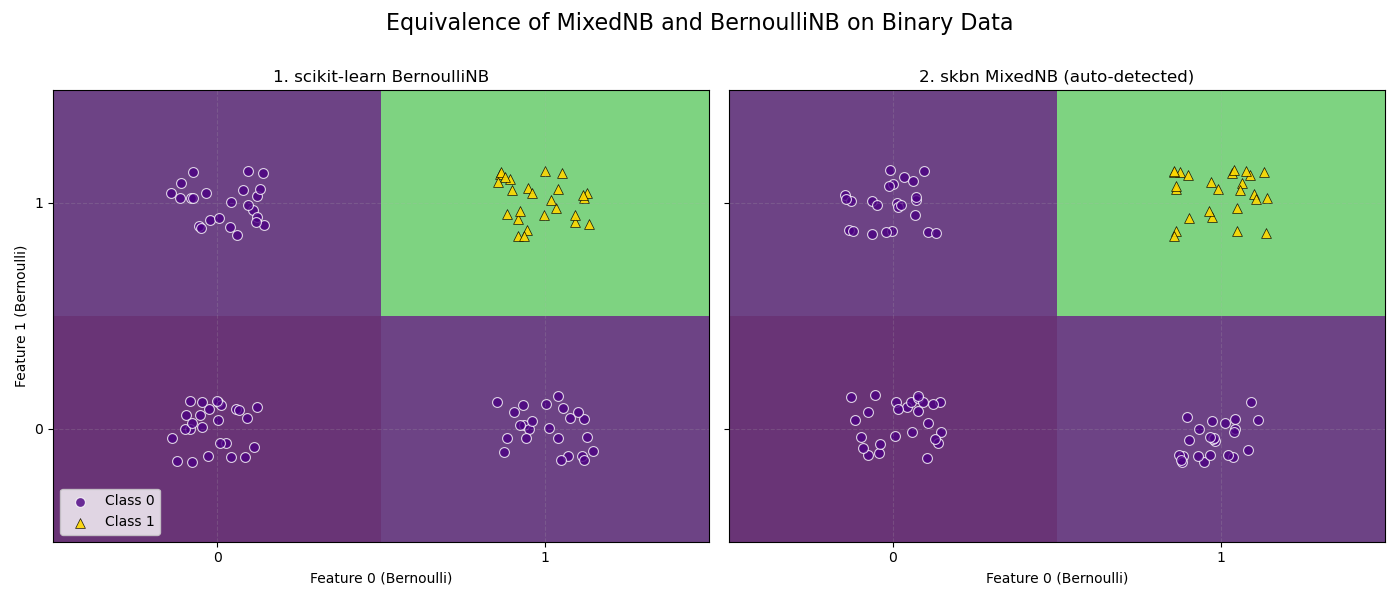
BernoulliNB vs MixedNB Equivalence Check: Probabilities are identical.
# Author: The scikit-bayes Developers
# SPDX-License-Identifier: BSD-3-Clause
import matplotlib.pyplot as plt
import numpy as np
from numpy.testing import assert_allclose
from sklearn.naive_bayes import BernoulliNB
from skbn.mixed_nb import MixedNB
# 1. Generate a 2D Bernoulli dataset (0s and 1s)
np.random.seed(42)
n_samples = 100
X = np.random.randint(0, 2, size=(n_samples, 2), dtype=int)
# Logic: AND gate (Class 1 only if both are 1)
y = (X[:, 0] & X[:, 1]).astype(int)
# 2. Fit both classifiers
bnb = BernoulliNB(alpha=1.0)
bnb.fit(X, y)
probs_bnb = bnb.predict_proba(X)
# MixedNB will auto-detect both features as 'bernoulli' (unique_values=2)
mnb = MixedNB(alpha=1.0)
mnb.fit(X, y)
probs_mnb = mnb.predict_proba(X)
# 3. Assert equivalence
try:
assert_allclose(probs_bnb, probs_mnb, rtol=1e-7, atol=1e-7)
equivalence_message = "Probabilities are identical."
except AssertionError as e:
equivalence_message = f"Probabilities are NOT identical:\n{e}"
print(f"BernoulliNB vs MixedNB Equivalence Check: {equivalence_message}")
# 4. Plot decision boundaries
fig, axes = plt.subplots(1, 2, figsize=(14, 6), sharey=True)
models = [bnb, mnb]
titles = ["1. scikit-learn BernoulliNB", "2. skbn MixedNB (auto-detected)"]
# Define grid (Centers for prediction, Edges for plotting)
# Binary features: 0, 1
centers = np.array([0, 1])
edges = np.array([-0.5, 0.5, 1.5])
# Create prediction grid
xx, yy = np.meshgrid(centers, centers)
grid_pred = np.c_[xx.ravel(), yy.ravel()]
for ax, model, title in zip(axes, models, titles):
# Predict probabilities on binary grid
probs = model.predict_proba(grid_pred)[:, 1]
Z = probs.reshape(xx.shape)
# Plot Heatmap - VIRIDIS
# Using pcolormesh with edges defines the 4 quadrants perfectly
ax.pcolormesh(
edges,
edges,
Z,
cmap="viridis",
vmin=0,
vmax=1,
shading="flat",
alpha=0.8,
edgecolors="none",
)
# Overlay real data points with consistent style
# Jitter points slightly to show density (since many points overlap on 0/1)
x_jit = X[:, 0] + np.random.uniform(-0.15, 0.15, size=n_samples)
y_jit = X[:, 1] + np.random.uniform(-0.15, 0.15, size=n_samples)
# Class 0 -> Indigo Circle
ax.scatter(
x_jit[y == 0],
y_jit[y == 0],
c="indigo",
marker="o",
s=50,
alpha=0.8,
edgecolors="w",
linewidth=0.8,
label="Class 0",
)
# Class 1 -> Gold Triangle
ax.scatter(
x_jit[y == 1],
y_jit[y == 1],
c="gold",
marker="^",
s=50,
alpha=0.9,
edgecolors="k",
linewidth=0.5,
label="Class 1",
)
ax.set_title(title, fontsize=12)
ax.set_xticks([0, 1])
ax.set_yticks([0, 1])
ax.grid(True, alpha=0.2, linestyle="--")
axes[0].set_ylabel("Feature 1 (Bernoulli)")
axes[0].set_xlabel("Feature 0 (Bernoulli)")
axes[1].set_xlabel("Feature 0 (Bernoulli)")
axes[0].legend(loc="lower left")
fig.suptitle("Equivalence of MixedNB and BernoulliNB on Binary Data", fontsize=16)
plt.tight_layout()
plt.subplots_adjust(top=0.85)
plt.show()
Total running time of the script: (0 minutes 0.128 seconds)
