Note
Go to the end to download the full example code.
Handling Mixed Data Types with MixedNB#
This example compares three strategies for handling a dataset with mixed continuous (Gaussian) and discrete (Categorical) features.
The Scenario (Generative Data):
We generate data from two natural clusters (Classes 0 and 1):
Continuous Feature: Two overlapping Gaussian distributions. Precision is key here.
Categorical Feature: Different category probabilities per class. Class 0 prefers Cat ‘0’, Class 1 prefers Cat ‘2’.
The Competitors:
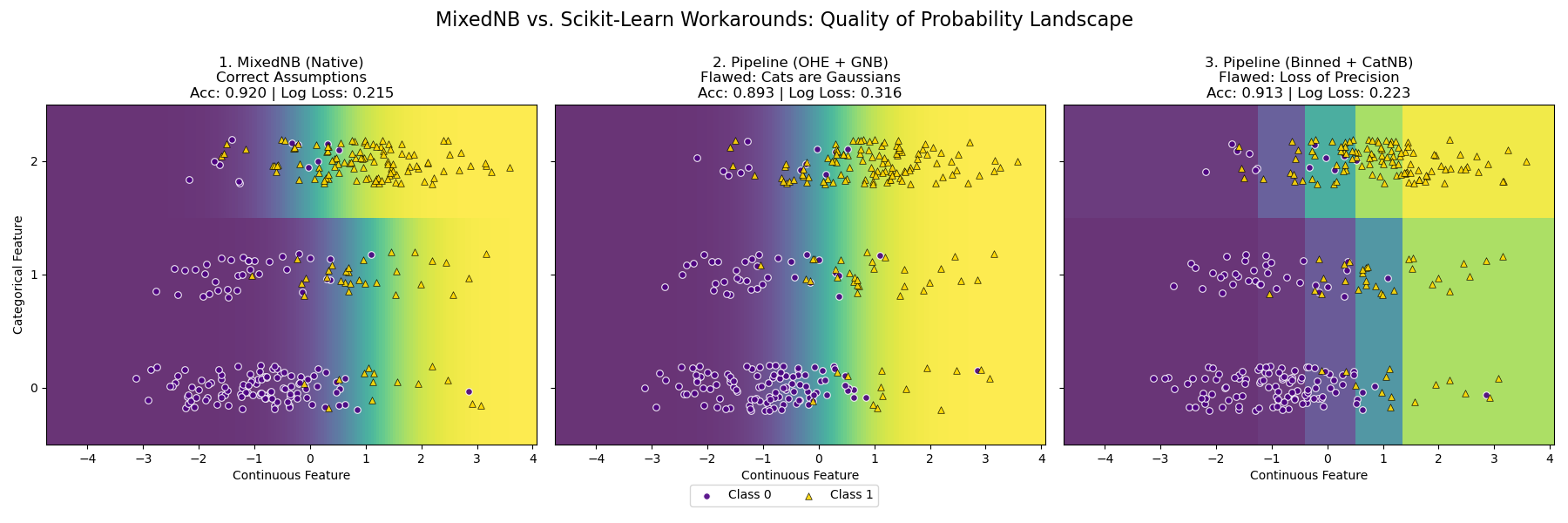
MixedNB (Native): Models Gaussian as Gaussian, Categorical as Multinomial. Matches the data generation process. (Acc: ~0.920).
Pipeline (OHE + GNB): Treats categories as binary Gaussians. (Acc: ~0.893).
Pipeline (Discretizer + CatNB): Bins continuous data. (Acc: ~0.913).
Result:
MixedNB produces the smoothest probability landscape and optimal log-loss, though standard pipelines perform reasonably well on this simple problem.
# Author: The scikit-bayes Developers
# SPDX-License-Identifier: BSD-3-Clause
import warnings
import matplotlib.pyplot as plt
import numpy as np
from sklearn.compose import ColumnTransformer
from sklearn.metrics import accuracy_score, log_loss
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import CategoricalNB, GaussianNB
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import KBinsDiscretizer, OneHotEncoder
from skbn.mixed_nb import MixedNB
# Suppress warnings
warnings.filterwarnings("ignore")
# --- 1. Generate Probabilistic Mixed Data ---
np.random.seed(42)
n_samples = 1000
# Class 0: Centered at X=-1, Prefer Cat 0
n0 = n_samples // 2
X_cont_0 = np.random.normal(-1.0, 1.0, size=n0)
# Probabilities for Cat 0, 1, 2: [0.7, 0.2, 0.1]
X_cat_0 = np.random.choice([0, 1, 2], size=n0, p=[0.7, 0.2, 0.1])
# Class 1: Centered at X=1, Prefer Cat 2
n1 = n_samples - n0
X_cont_1 = np.random.normal(1.0, 1.0, size=n1)
# Probabilities for Cat 0, 1, 2: [0.1, 0.2, 0.7]
X_cat_1 = np.random.choice([0, 1, 2], size=n1, p=[0.1, 0.2, 0.7])
# Combine
X_cont = np.concatenate([X_cont_0, X_cont_1])
X_cat = np.concatenate([X_cat_0, X_cat_1])
y = np.concatenate([np.zeros(n0), np.ones(n1)]).astype(int)
# Stack: [Continuous, Categorical]
X = np.column_stack([X_cont, X_cat])
# Split for valid metric calculation
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=42
)
# --- 2. Define Models ---
# Model 1: MixedNB
mnb = MixedNB()
mnb.fit(X_train, y_train)
# Model 2: OHE + GaussianNB
pipe_ohe = make_pipeline(
ColumnTransformer(
[("onehot", OneHotEncoder(handle_unknown="ignore", sparse_output=False), [1])],
remainder="passthrough",
),
GaussianNB(),
)
pipe_ohe.fit(X_train, y_train)
# Model 3: Discretizer + CategoricalNB
pipe_kbins = make_pipeline(
ColumnTransformer(
[
(
"discretizer",
KBinsDiscretizer(n_bins=5, encode="ordinal", strategy="quantile"),
[0],
)
],
remainder="passthrough",
),
CategoricalNB(),
)
pipe_kbins.fit(X_train, y_train)
models = [mnb, pipe_ohe, pipe_kbins]
titles = [
"1. MixedNB (Native)\nCorrect Assumptions",
"2. Pipeline (OHE + GNB)\nFlawed: Cats are Gaussians",
"3. Pipeline (Binned + CatNB)\nFlawed: Loss of Precision",
]
# --- 3. Visualization ---
fig, axes = plt.subplots(1, 3, figsize=(18, 6), sharey=True)
# Grid for plotting
h = 0.05
x_min, x_max = X[:, 0].min() - 0.5, X[:, 0].max() + 0.5
y_min, y_max = -0.5, 2.5 # Categories 0, 1, 2
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, 1))
# Flatten for prediction
grid_X = np.c_[xx.ravel(), yy.ravel()]
for ax, model, title in zip(axes, models, titles):
# Predict
Z = model.predict_proba(grid_X)[:, 1]
Z = Z.reshape(xx.shape)
# Metrics on Test Set
acc = accuracy_score(y_test, model.predict(X_test))
ll = log_loss(y_test, model.predict_proba(X_test))
# Plot Heatmap - VIRIDIS
# 0.0 = Purple (Class 0 zone), 1.0 = Yellow (Class 1 zone)
ax.imshow(
Z,
extent=(x_min, x_max, y_min, y_max),
origin="lower",
cmap="viridis",
vmin=0,
vmax=1,
aspect="auto",
alpha=0.8,
)
# Overlay real data points (with jitter on Y)
X_plot = X_test.copy()
y_jit = X_plot[:, 1] + np.random.uniform(-0.2, 0.2, size=len(X_plot))
# Visual Coherence with Viridis:
# Class 0 -> Indigo (Matches background purple)
# Class 1 -> Gold (Matches background yellow)
# Edgecolors='w' ensures visibility even on matching backgrounds
ax.scatter(
X_plot[y_test == 0, 0],
y_jit[y_test == 0],
c="indigo",
marker="o",
s=30,
alpha=0.9,
edgecolors="w",
linewidth=0.8,
label="Class 0",
)
ax.scatter(
X_plot[y_test == 1, 0],
y_jit[y_test == 1],
c="gold",
marker="^",
s=30,
alpha=0.9,
edgecolors="k",
linewidth=0.5,
label="Class 1",
) # Black edge for yellow points for better contrast
# Title & Metrics
ax.set_title(f"{title}\nAcc: {acc:.3f} | Log Loss: {ll:.3f}", fontsize=12)
ax.set_xlabel("Continuous Feature")
ax.set_yticks([0, 1, 2])
axes[0].set_ylabel("Categorical Feature")
# Clean legend
handles, labels = axes[0].get_legend_handles_labels()
fig.legend(handles, labels, loc="lower center", ncol=2, bbox_to_anchor=(0.5, 0.02))
fig.suptitle(
"MixedNB vs. Scikit-Learn Workarounds: Quality of Probability Landscape",
fontsize=16,
)
plt.tight_layout()
plt.subplots_adjust(top=0.80, bottom=0.15)
plt.show()
Total running time of the script: (0 minutes 0.284 seconds)
